AROMMA
Paper: https://doi.org/10.48550/arXiv.2601.19561
Code: https://github.com/DGIST-Distributed-AI-Lab/aromma
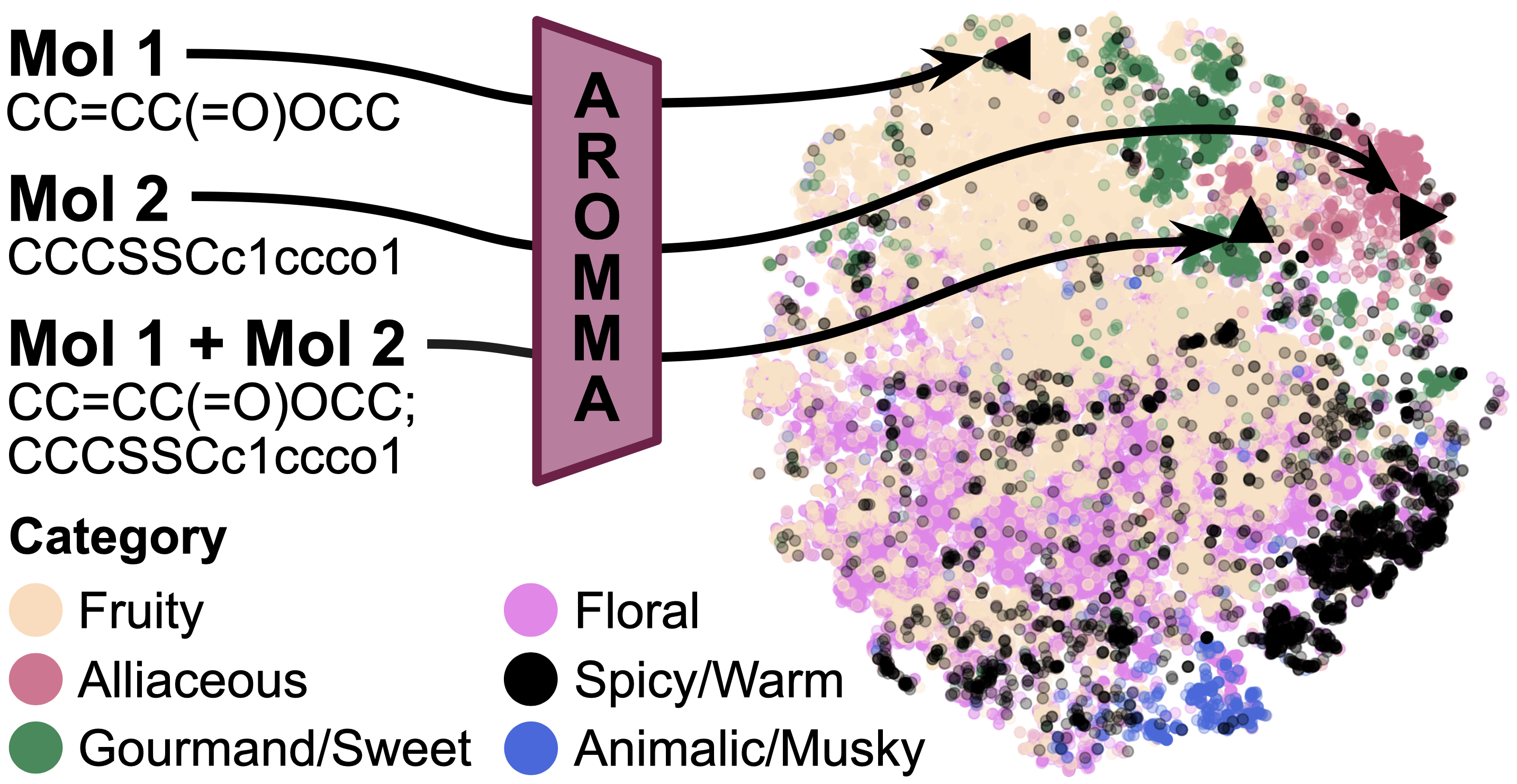
AROMMA is a novel framework that learns a unified embedding space for both single molecules and binary mixtures, providing a generalized representation for olfactory perception.
The framework leverages a large-scale chemical foundation model (e.g., SPMM) for robust molecular representations, incorporates an attention-based aggregator to model permutation invariance and asymmetric interactions, and addresses label sparsity through knowledge distillation and pseudo-labeling.
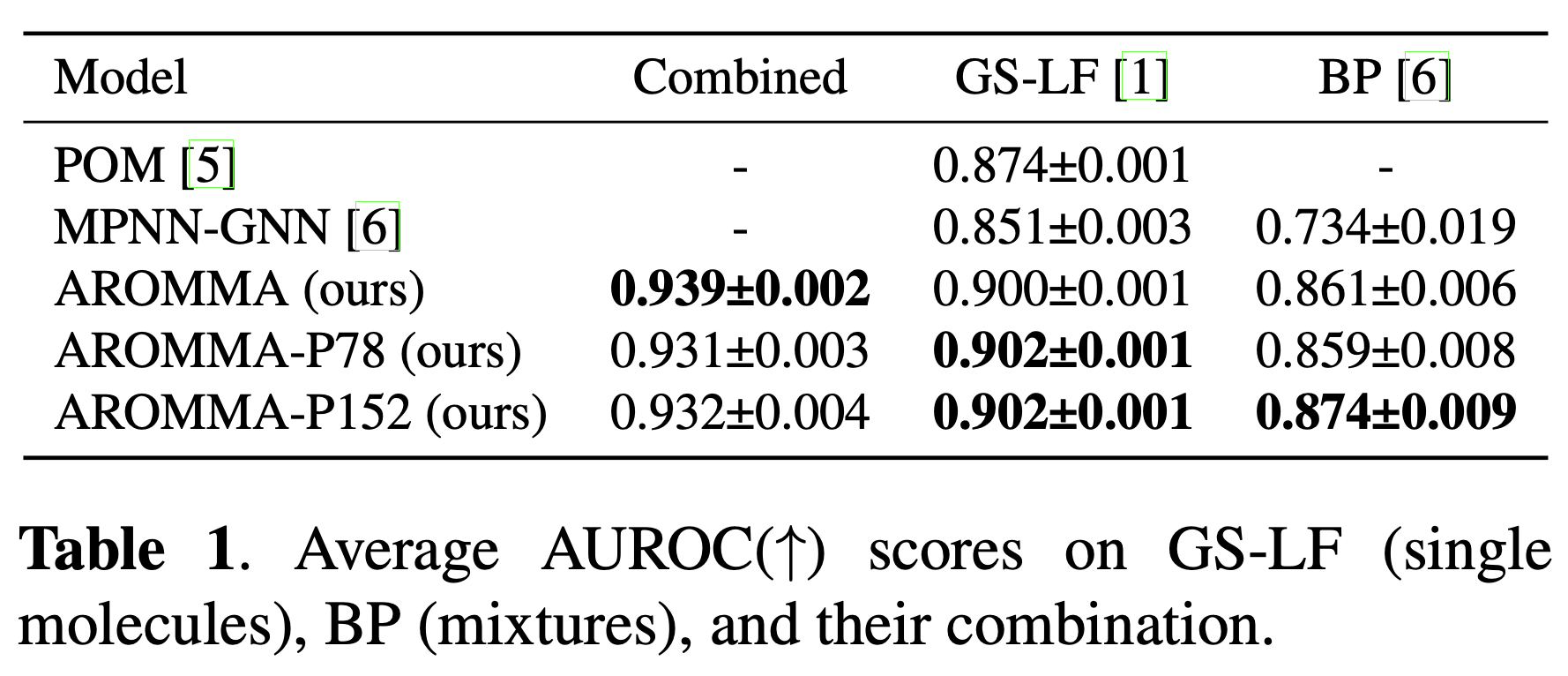
AROMMA achieves state-of-the-art performance on both GS-LF (single-molecule) and BP (binary-mixture) benchmarks, with up to 19.1% improvement in AUROC, demonstrating effective bidirectional knowledge transfer and strong generalization ability.
Architecture Overview
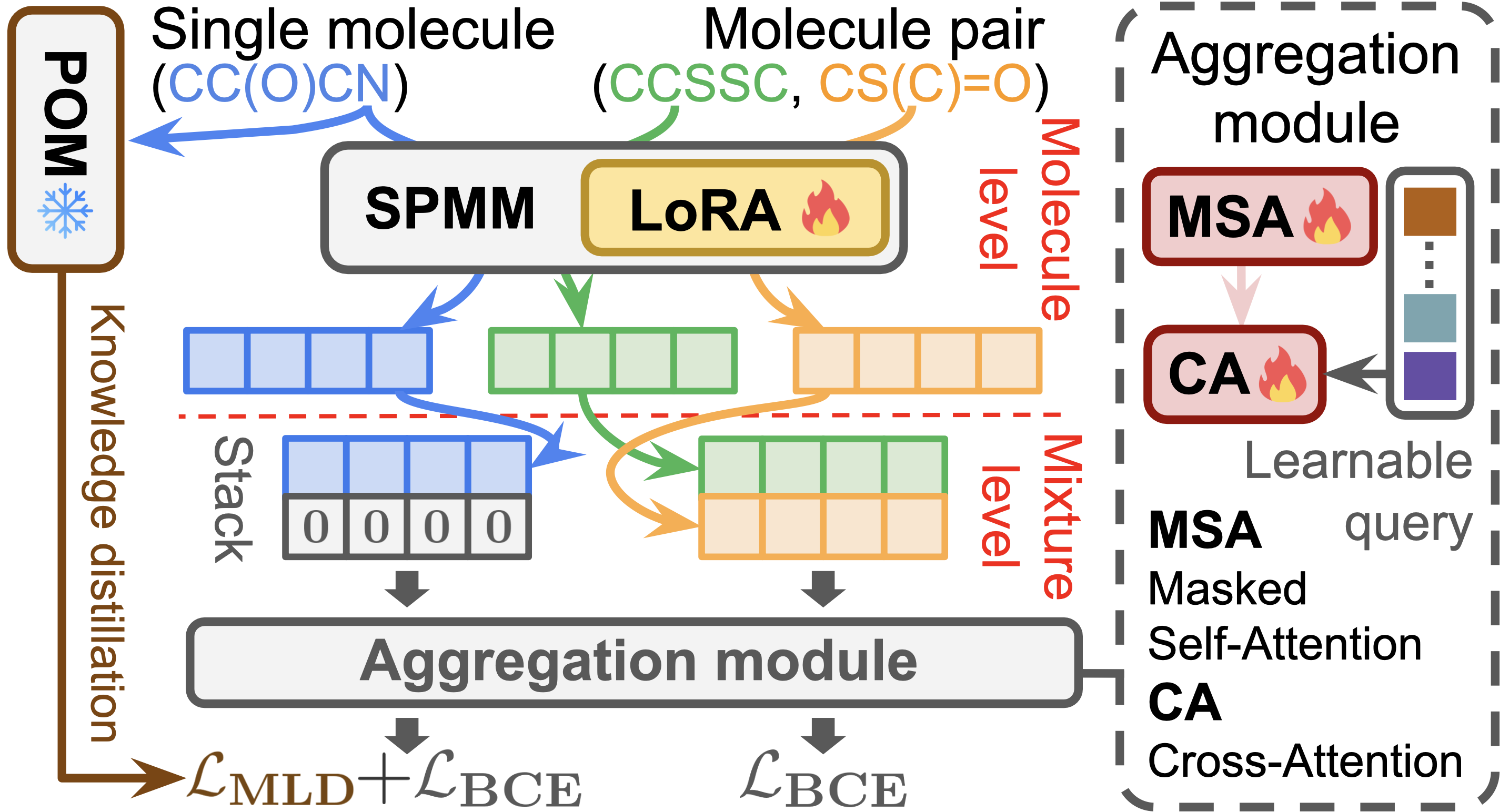
Embedder
- Individual molecular embeddings are obtained using the SMILES encoder of SPMM.
- SPMM is pre-trained on approximately 50 million molecules, providing robust molecule representations.
- The model is fine-tuned efficiently using LoRA (Low-Rank Adaptation).
Aggregation Module
- The input consists of SPMM embeddings \(E \in \mathbb{R}^{2\times d_e}\) (zero-padded in the case of a single molecule).
- The embeddings pass through a linear projection followed by a ReLU activation, producing \(E'\).
- A masked multi-head self-attention layer generates \(H\), which represents contextualized molecular embeddings.
- The contextualized representations \(H\) are used as keys and values in a cross-attention layer, together with a global learnable query \(q \in \mathbb{R}^{1 \times d'_h}\).
- Finally, Layer Normalization is applied to obtain the global embedding \(z\).
Training Strategy
Loss Function
- Initial training is conducted on the unified 152-descriptor set, where the 78 missing BP labels are treated as zeros.
- For single molecules, we combine:
- Multi-Label Logit Distillation (MLD) loss for knowledge distillation: POM, a state-of-the-art model for single-molecule odor prediction, serves as the teacher to alleviate label sparsity \(\rightarrow \mathcal{L}_{MLD}(P^S,P^T)=\mathcal{D}_{KL}(P^T||P^S)+\mathcal{D}_{KL}((1-P^T)||(1-P^S))\)
- Binary Cross-Entropy (BCE) loss for supervised learning
- For binary mixtures, only the BCE loss is used.
The total loss balances single-molecule and mixture objectives with loss balancing coefficient.
Latent Label Augmentation & Re-training
To address label sparsity and incompleteness in the BP dataset, we apply class-aware pseudo-labeling.
- We first compute the positive label ratio for each class \(c\) in GS-LF \(\gamma_c = \frac{1}{n} \sum_{j=1}^n \mathbf{1}(y_{jc}=1)\).
- For each class \(c\), we select a threshold \(\tau_c\) such that the proportion of predicted probabilities exceeding the threshold matches the class prior \(\frac{1}{n} \sum_{j=1}^n \mathbf{1}(p_{jc} \ge \tau_c)=\gamma_c\).
- Using this strategy, we construct two augmented datasets Pseudo-78 and Pseudo-152. The model is then re-trained as AROMMA-P78 and AROMMA-P152.
Results
- Extended annotations include more fine-grained odor descriptors. For instance, blending
OC1COC(Cc2ccccc2)OC1withOCc1ccccc1yields ground-truth labels such as floral and fruity, while our model additionally identifies rose, a descriptor within the floral category. - Our method achieves state-of-the-art performance on both GS-LF and BP, with pseudo-labeled data further boosting performance, highlighting the effectiveness of our approach.